This work was carried out in joint cooperation between two active Oil and Gas operators.
This is our second completed pilot publication, executed just after our Argentina pilot. We’re extremely excited to share these results too. This pilot was performed in the USA; Biodentify applied and blind tested patented DNA fingerprinting technology and, using advanced machine learning algorithms, accurately predicted hydrocarbon presence in the subsurface prior to drilling.
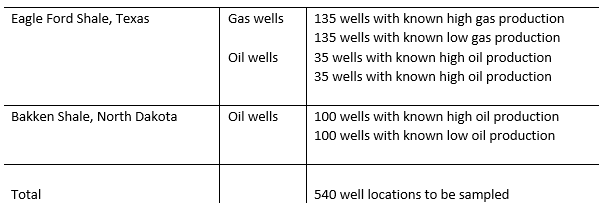
The Eagle Ford Shale Formation of Southern Texas was chosen as both an unconventional oil and unconventional gas play, whereas the Bakken Shale Formation of North Dakota was chosen as a pure unconventional oil play. Both of these plays have seen substantial exploration and production activities over the past 15-20 years. This project delivered proof of technology in the form of a highly accurate map of subsurface hydrocarbon productivity, predicted with 85% accuracy at 200 blinded sample locations. All results are based on real data from our own fieldwork and we also ranked results, distinguishing high versus low producing wells.
Biodentify’s patented technology is based on the response elicited in the shallow soil (<30cm) microbial population by hydrocarbons which have migrated from depth to the surface through vertical microseepage. The trace hydrocarbons released are otherwise undetectable, but the bacterial population acts as an extremely sensitive indicator. Samples are collected and analyzed for microbial DNA content leading to a characteristic fingerprint, from which final predictions can be made.
Planning the project out with our customer
In 2018, Biodentify was awarded a research grant by the EU Horizon H2020 fund. This EU grant was used to set up research pilots in three different continents, and the second continent we worked on was the USA.
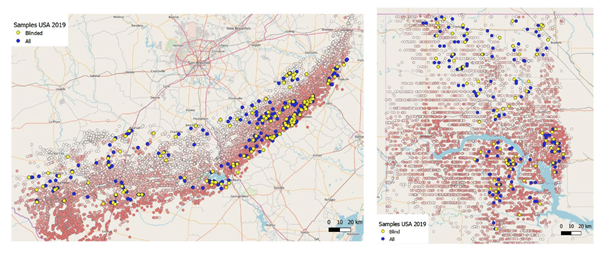
We concentrated on sampling locations with known production statistics, taking a total of 540 samples over 34,000 km. Three different sets of samples were taken: high and low producing wells in the Bakken (oil shale in North Dakota) and high and low producing wells in both the oil window and the gas window in the Eagle Ford (oil and gas shale in Texas). For validation, 200 known samples were blinded and relabeled by our customer after the sample taking (70 from the Bakken, 30 from the Eagle Ford oil window and 100 from the Eagle Ford gas window). This was in order for us to be able to once again validate our predictions.
Our USA pilot methodology
We began executing our research in an area of 310 by 65 km (Texas) and 160 by 80 km (North Dakota). In these areas, the top percentile from both the highest and lowest producing wells was selected. These wells were ranked based on normalized data, taking into account and corrected for their vintage and lateral length. Once wells were chosen, the surface intersection of the lateral area was sampled and we covered the vast areas relatively quickly with a field crew and car. Our predictions were again made using only shallow soil (<30cm / 1 foot). Our patented technology is so sensitive and testing so effective, that we only need half a teaspoon of soil per sample.
We analyzed for microbes in our lab, separating the microbial DNA from the soil, ending up with a DNA fingerprint for each location. This data is very vast, hundreds of millions of numerical values, too vast for us humans to make sense of it alone, which is where supercomputing power comes in. The use of our unique and constantly updated DNA database, alongside big data algorithms, machine learning and our own metagenomic pipeline, ensure that the model is constantly enhanced and improved over time to enable us to accurately predict subsurface hydrocarbon presence.

Building a predictive model for the USA pilot using machine learning
There are two main steps in building a predictive model using machine learning:
1. Selecting a training set from our database with known productivity.
2. Modelling and validating the microbes that determine productivity.
Data source: ShaleProfile Analytics – cumulative production first two years, normalized for vintage and lat length. After this the cumulative production was normalized, on the ranking per calendar year (vs whole production period) and on the production divided by lateral length. This results in four criteria or scores between 0-1 :
- 2 years cumulative production over the entire period of production
- 2 years cumulative production per calendar year
- 2 years cumulative production per lateral length
- 2 years cumulative production per lateral length per year
The 248 successful non-blinded DNA fingerprints were used to train the model. Machine learning uses a strict protocol to check if the dataset is ‘trainable’, meaning that there is a correlation found that reflects a predictive model. In modelling terms this is: that the model is not overfitted and only reflects detail that fits the training dataset—and will not accurately predict new datasets. Our training sets are generated by randomly selecting 70% of these samples and validating the best predictive model on the 30% left out. Repeating this procedure a thousand times delivers the basis of estimates and cross-validation statistics. This modelling approach prevents overfitting and gets the highest prediction accuracy, and we executed it through a two stage approach: i) predict which of the 143 blinded samples were from the Eagle Ford and the Bakken respectively and ii) train separate models for non-blinded Eagle Ford and Bakken to predict the subsets belonging to step 1.
Uncovering of the blinded samples by our customer
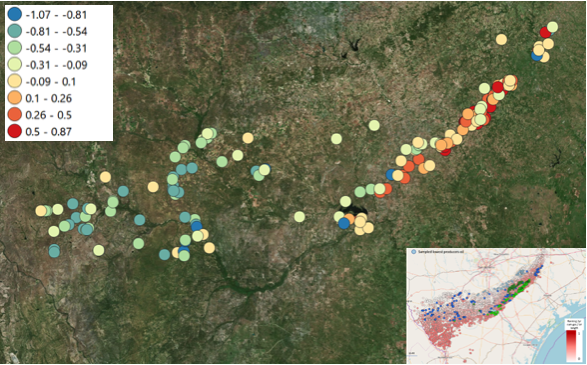
Building and validating our training set, and with that creating a robust prediction model, meant that some 2000 models had to be run in order to predict the truly blinded samples. Results of these predictions turned out to be 85% accurate, after our customer uncovered them. It showed that the (internal) cross validation of our training models had nearly the same accuracy as the predictions of the blinded samples, meaning that the models were robust and not overfitted.
Exportability of the models we tested
We also tested the ‘exportability’ of our trained models. It became apparent our first models, only trained on the Bakken data, could also be used to predict the Eagle Ford samples. This could be done with an accuracy of 72%. It demonstrates that our trained models are exportable over larger distances and to other plays with different geography, geology and hydrocarbon settings.
This exportability was further confirmed by using the trained models for both US plays to predict locations in the Vaca Muerta of Argentina (described here) where Biodentify obtained an accuracy of 83%.
In both cases the accuracy dropped when compared with using models based on local data, which makes sense, but also shows there is an overlap in the selected microbes acting as an indicator of oil and gas. This aspect has to be investigated further to draw more detailed and general conclusions.
USA pilot results and conclusions
A) The large number of shale wells in the US make it possible to design a setup for training a machine learning model with a high number of locations where hydrocarbon production from shale wells is relatively low or high, and that these two can be distinguished from each other.
B) We concluded that distinguishing between sample locations above low producing wells versus sample locations above high producing wells for this dataset in two distinctively different shale plays in the US is possible.
C) Having local samples increases the accuracy but we can also export our models: our trained model on Bakken data was used to predict the Eagle Ford samples and a high accuracy was achieved. This means that our trained models are exportable over large distances to other plays with different geography, geology and hydrocarbon settings.
Additionally this exportability was further confirmed by using the trained models for both US plays to predict locations in the Vaca Muerta, Argentina, our first pilot project.
Our main conclusion is that our methodology does, before drilling and without knowing anything about the subsurface conditions, very accurately predict the presence of hydrocarbons in the subsurface. Secondly, we successfully used a model from one continent and accurately predicted a second hydrocarbon area over 8000 km away. This gives us the huge potential to use our uniquely calibrated database further in other new locations, as well as the improving accuracy/usability of our DNA fingerprinting model and database.
We believe these results are sufficiently interesting for major oil and gas companies to apply this technology in their own projects, minimizing risk, maximizing returns and saving money.
To find out more, do not hesitate to get in touch here.
About Biodentify
Biodentify was founded as a spin-out from Dutch R&D group TNO in December 2014. TNO is a 3000+ R&D organization and the largest microbiological research group in the Netherlands. Biodentify is owned and managed by 3 partners with extensive entrepreneurial experience bringing innovative new technologies to market in the Oil & Gas industry. The company has developed a novel and patented technology, predicting prospectivity before drilling, with > 70% accuracy, based on microbial DNA analysis of shallow soil or seabed samples through unique machine learning models. Biodentify was named the #58 most innovative company in 2019 by the KvK (Chamber of Commerce).